Adds ghost convert_to_mat — convert ENVI, TIFF, GeoTIFF, and HDF5/NetCDF hyperspectral images to .mat format. Auto-detects format from extension, preserves all metadata (CRS, wavelengths, band names) in a JSON sidecar, and supports ground truth in .mat, .png, .tif, or .hdr.
pip install ghost-hsi[convert]
ghost convert_to_mat \
--img image.hdr \
--gt labels.tif \
--out-dir converted/What does "generalizable" actually mean here?
For a hyperspectral model to work across different scenes without retraining, a few things need to be true. Here's where GHOST currently stands.
Band count, class count, and spatial dimensions are all read from the file at runtime. Nothing is hardcoded. Same binary on Indian Pines, LUSC, and Mars CRISM.
Works on 3 bands or 400+. Same pipeline, any sensor resolution, no configuration needed.
Remote sensing, medical pathology, planetary science — no format-specific code paths, no domain-specific preprocessing.
For true scene-to-scene transfer, the model needs to classify each pixel from its spectrum alone — no spatial context from neighbors. We're not there yet.
Results.
Tested on standard HSI benchmarks. These numbers are from single-scene pixel-level evaluation, which is the standard for these datasets but doesn't say much about real-world generalization. Expect roughly ±1% variance between runs due to random splits and seed sensitivity. Take them accordingly.
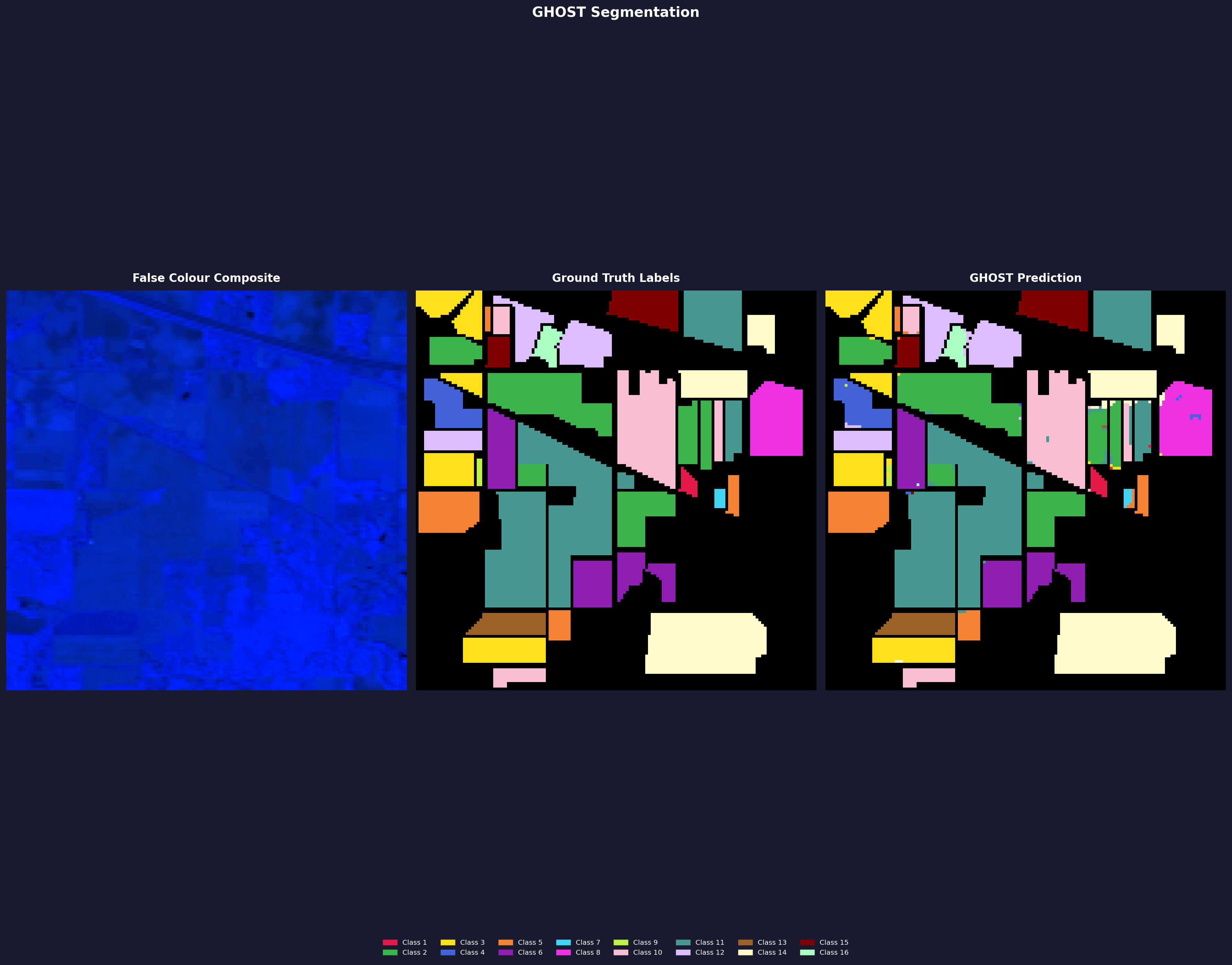
Indian Pines
Metrics
| Config | OA (%) | mIoU | Kappa |
|---|---|---|---|
| 64 / 16 | 98.16 | 0.9071 | 0.9790 |
| 32 / 8 | 97.20 | 0.8030 | 0.9681 |
Config
| Base / Num Filters | GPU | Time |
|---|---|---|
| 64 / 16 | RTX 3050 (laptop) | 2h 20m |
| 32 / 8 | RTX 3050 (laptop) | 1h 17m |
Standard pixel-level split — 20% train, 10% val, 70% test, seed 42, forest routing, dice loss. The image shown is the 64/16 run. Both runs use the same scene — evaluation is inflated relative to cross-scene generalization.
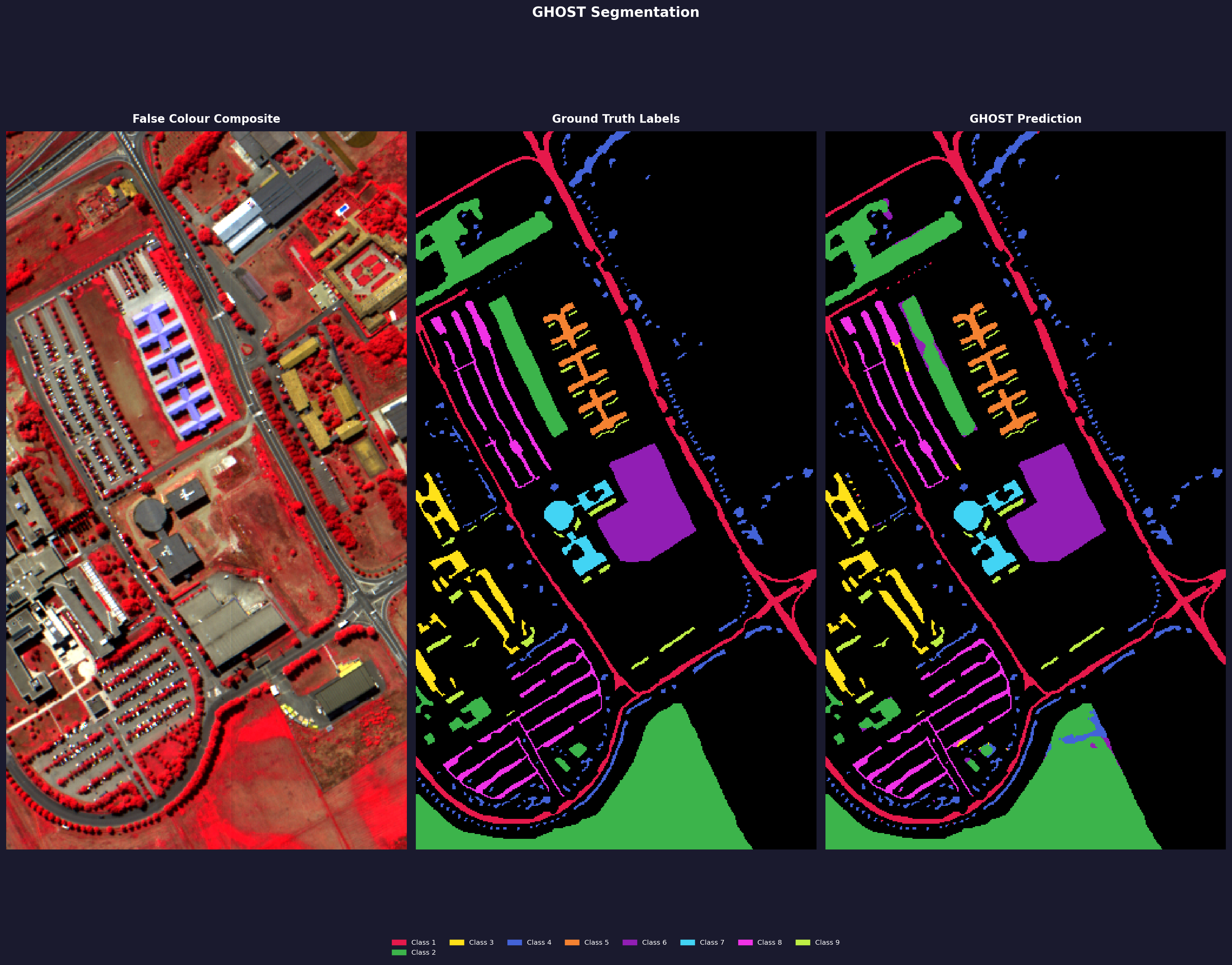
Pavia University
Metrics
| Config | OA (%) | mIoU | Kappa |
|---|---|---|---|
| 32 / 8 | 97.47 | 0.9531 | 0.9667 |
Config
| Base / Num Filters | GPU | Time |
|---|---|---|
| 32 / 8 | Kaggle T4 | 7h 29m |
Standard pixel-level split, seed 42, forest routing, dice loss. Ran on Kaggle free tier T4.
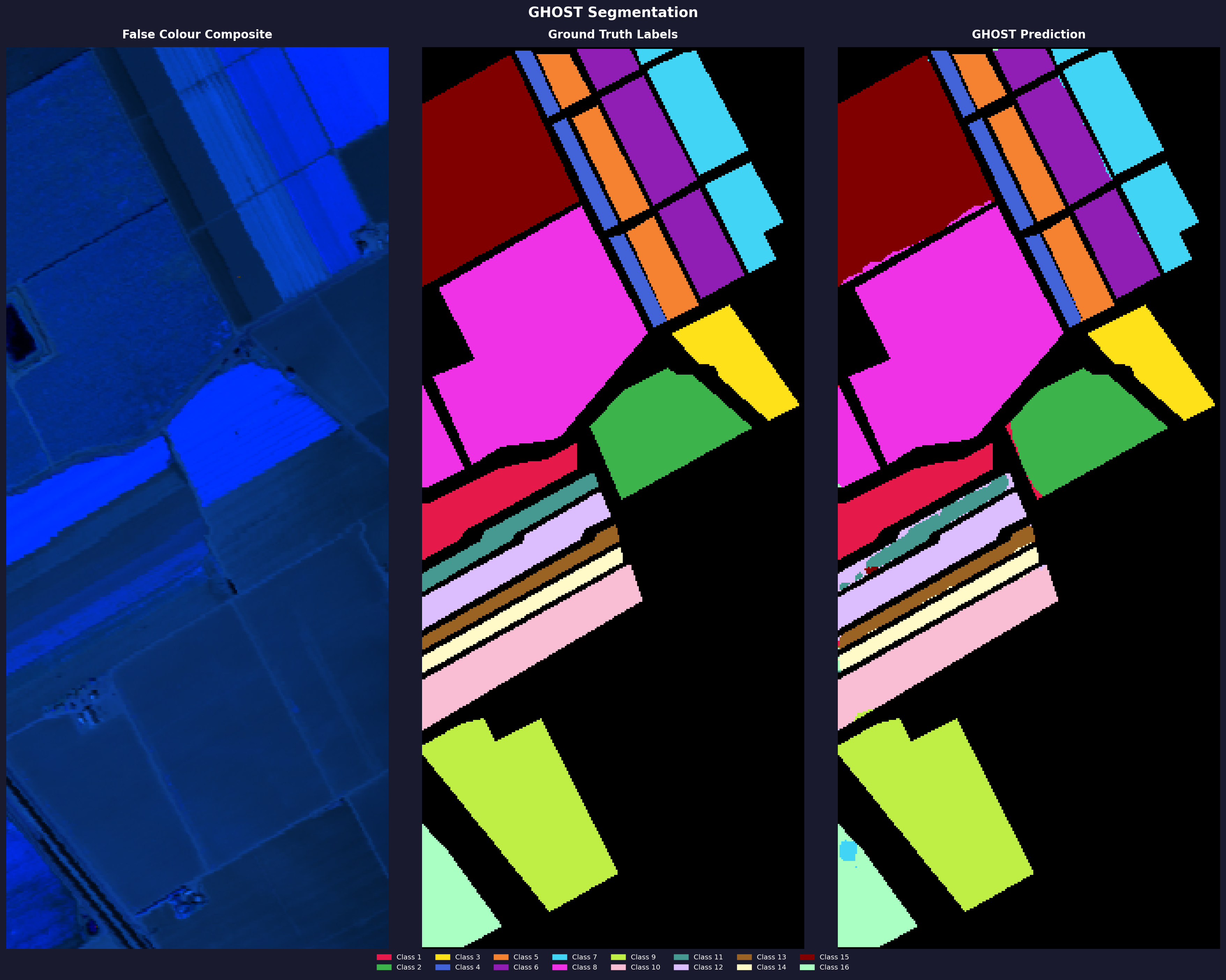
Salinas Valley
Metrics
| Config | OA (%) | mIoU | Kappa |
|---|---|---|---|
| 32 / 8 | 98.69 | 0.9577 | 0.9855 |
Config
| Base / Num Filters | GPU | Time |
|---|---|---|
| 32 / 8 | Kaggle T4 | 10h 51m |
Standard pixel-level split, seed 42, forest routing, dice loss. Ran on Kaggle free tier T4.
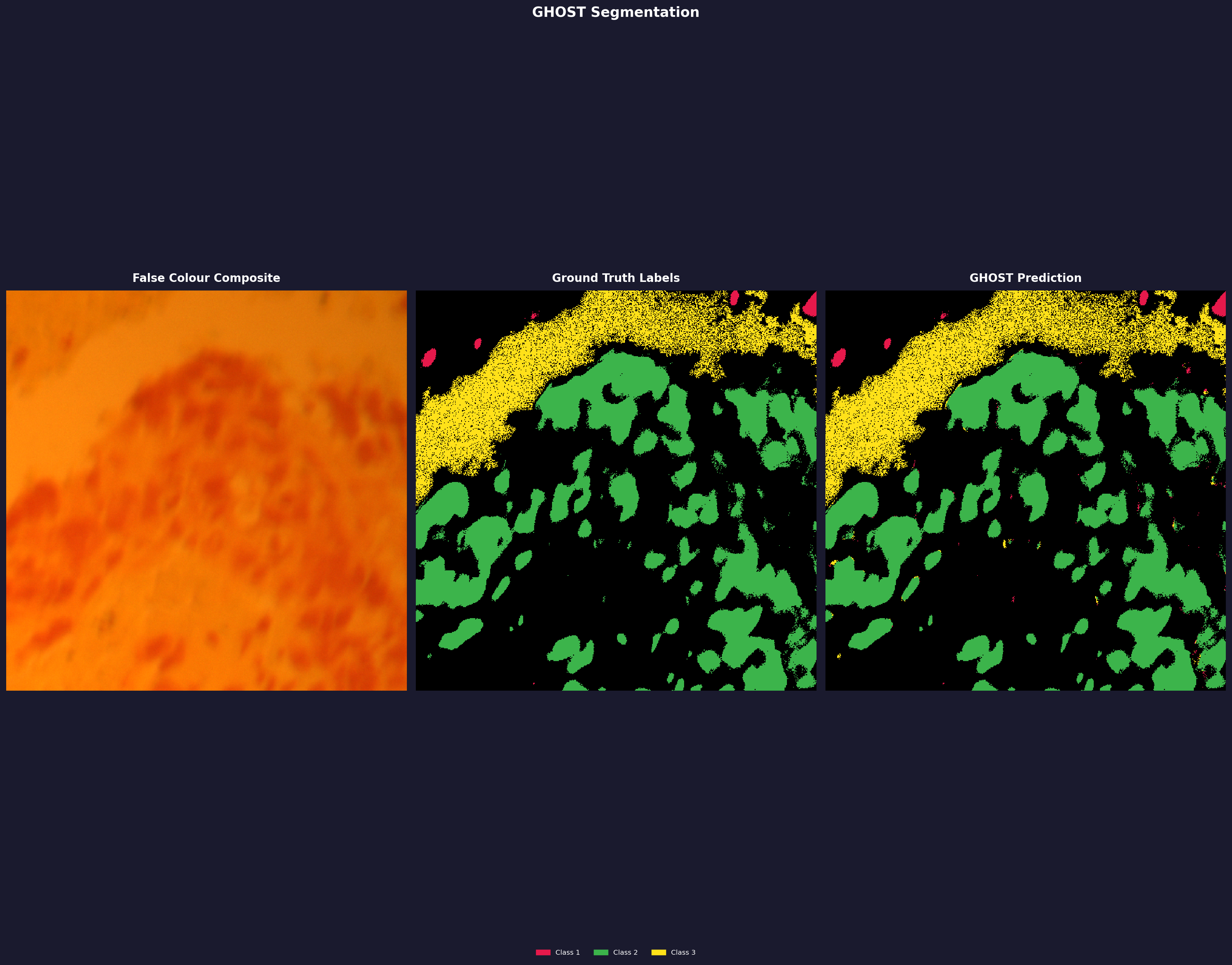
LUSC (Lung Cancer)
Metrics
| Config | OA (%) | mIoU | Kappa |
|---|---|---|---|
| 32 / 8 | 99.42 | 0.9263 | 0.9876 |
Config
| Base / Num Filters | GPU | Time |
|---|---|---|
| 32 / 8 | RTX 3050 (laptop) | 1h 8m |
Single tissue result only. This is from one 512×512 crop from a single patient scan — it is not representative of the full LUSC dataset and cannot be compared to published clinical benchmarks. The dataset has 60+ scans; multi-scan evaluation is a v0.2.x goal. Dataset: HMI Lung Dataset.
Mars CRISM
Metrics
| Config | OA (%) | mIoU | Kappa |
|---|---|---|---|
| 32 / 8 | 71.70 | 0.5228 | 0.6829 |
Config
| Base / Num Filters | GPU | Time |
|---|---|---|
| 32 / 8 | Kaggle T4 | 6h 44m |
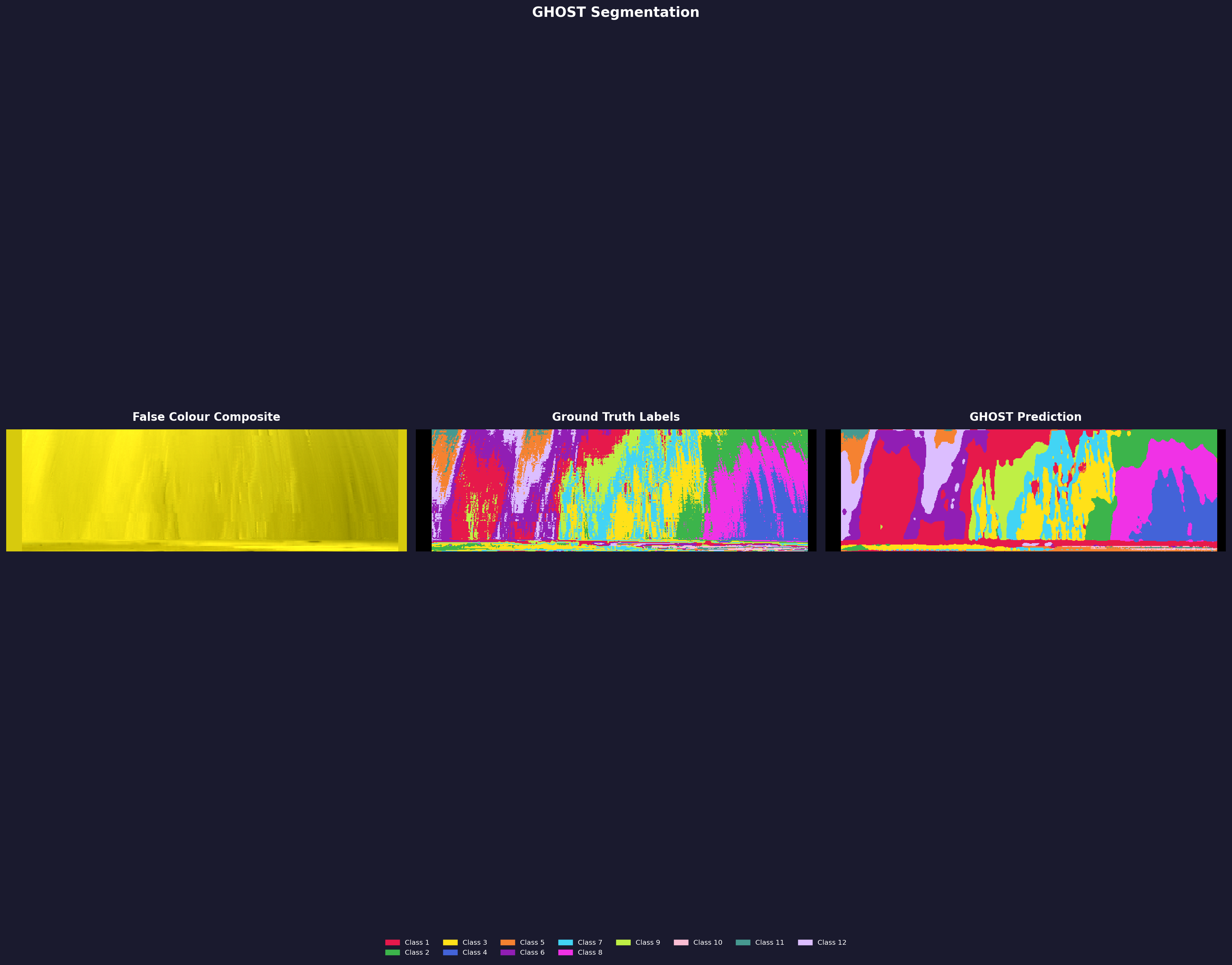
Quantitative results pending a clean run. The CRISM ground truth annotations are extremely sparse and noisy — the labels themselves are grainy, which drags down the numbers even when GHOST produces a visually smooth and consistent segmentation. The image shows what GHOST predicted. Annotation quality is a known limitation of this dataset. More details once I have a proper evaluation set up. Dataset: NASA MRO/CRISM.
Why no comparisions ?
Every published HSI method I'm aware of — including v0.1.x of this project — relies on spatial context. U-Nets, 3D CNNs, hybrid architectures: they all, to varying degrees, learn where things tend to be in a scene alongside what they look like spectrally. That's why they get high numbers on benchmarks. It's also why they don't generalize across scenes.
Comparing GHOST v0.1.x against those methods would be comparing like-for-like on a metric that doesn't test the actual thesis. The numbers might look competitive. They'd mean nothing.
The comparison that matters is this: does a spectral-only model, with no access to neighboring pixels at any stage, hold up across scenes it has never seen? That's the question v0.2.x is trying to answer. That's the benchmark worth publishing. Until that result exists — properly evaluated, with ablation studies, on multi-scene data — putting up a table of same-scene OA scores against the literature would just be adding more noise to a space that already has plenty of it.
Then why publish at all?
Because three of the four things GHOST set out to do are already done.
The same binary, unchanged, has segmented an AVIRIS remote sensing scene with 200 spectral bands, a lung cancer histopathology slide with 61 bands, and a Mars CRISM mineral map with ~400 bands. No format-specific code paths. No domain-specific preprocessing. No configuration changes between them. Data agnosticism, band-count agnosticism, sensor agnosticism — all three work.
Most HSI research treats remote sensing and medical imaging as entirely separate problems. GHOST doesn't. That's not nothing, even before cross-scene generalization is solved.
The fourth objective — true spectral-only generalization — is the hard one, and it isn't finished. But the foundation it needs is already there.
Architecture.
What's in the current build, and what's being redesigned.
Continuum Removal (Simplified)
A straight line is drawn from the first spectral band to the last, and the spectrum is divided by it. This removes the overall reflectance slope caused by illumination angle and atmospheric scatter — the dominant non-chemical variation in raw HSI data.
It's a reasonable starting point. The main limitation is that it doesn't normalize absorption features relative to their local context — a deep absorption dip sitting next to a tall reflectance peak and the same dip next to a short peak can look different in the output, even though the underlying chemistry is identical. Full convex hull continuum removal (planned for v0.2.x) handles this more properly.
Spectral 3D Conv Stack
Three sequential 3D convolutions with a (7, 3, 3) kernel — 7 bands deep spectrally, 3×3 spatially. The model learns that nearby spectral bands correlate with each other, and that nearby pixels tend to share materials.
The spatial component is both a strength and a problem. It captures local context, which helps within a single scene. But it also means the model is learning where things tend to be, not just what they look like spectrally. That doesn't transfer between scenes. This is one of the main things being removed in v0.2.x.
SE Attention Blocks
Squeeze-and-Excitation blocks learn a per-channel importance weight. A global average pool reduces each feature channel to a scalar, a small two-layer MLP predicts a weight between 0 and 1, and that weight is multiplied back into the channel. The network learns to suppress less useful channels and emphasize the ones that matter for the current input.
Lightweight and effective. No major issues with it — the spirit of per-channel weighting carries into v0.2.x implicitly through the dilated ResNet encoder.
U-Net Encoder–Decoder
A standard 4-level U-Net with skip connections. The encoder downsamples through MaxPool2d, the decoder upsamples via transposed convolutions, and skip connections link matching levels. Channel progression: f → 2f → 4f → 8f → 16f.
This is where most of the learning happens in v0.1.x — and also why the model doesn't generalize across scenes. The U-Net effectively learns "where things are" in the training image. Corn fields tend to appear in certain shapes, urban structures have spatial patterns. None of that is true in the next scene. That's the thing I'm trying to remove entirely in v0.2.x.
Spectral Partition Tree (SPT)
Previously called RSSP (Recursive Spectral Splitting with Parallel Forests), SPT is a divide-and-conquer strategy over the class set. It builds a binary tree where classes are recursively split based on SAM (Spectral Angle Mapper) distance — spectrally similar classes are grouped together and handled by specialist ensemble models.
The reasoning: asking one flat model to separate 16 highly imbalanced classes is hard. SPT lets one model split broad groups (vegetation vs. minerals), then separate specialists handle the hard within-group cases (corn subtypes, mineral variants). Each leaf ensemble focuses on a smaller, more tractable problem. In practice this helps a lot with class imbalance.
The SPT logic is unchanged in v0.2.x — only what lives inside each tree node changes.
SSSR Router (Experimental)
SSSR (Spectral State-Space Router) was an attempt to replace hard argmax routing between tree nodes with a learned spectral state-space model — differentiable, soft routing probabilities instead of hard decisions at each branch.
In practice it doesn't work. Ensemble routing (--routing forest) outperforms it in every configuration I've tested. The SSM pretraining adds significant wall-clock time, and the routing improvement simply doesn't materialize. The code is currently broken and not recommended.
It may be revisited in v0.1.7+ or rethought entirely for v0.2.x. For now, always use --routing forest and set --ssm_epochs 1 to skip SSM pretraining entirely.
Known Limitations
- Spatial dependence: The U-Net backbone processes neighboring pixels together. A model trained on one scene cannot reliably segment a different scene — it has learned spatial patterns alongside spectral ones.
- No transfer learning: Each new dataset requires a full training run from scratch. There's no way to reuse weights from a previous run in any meaningful way.
- Single scene only: Training and inference must be on the same scene, or at very minimum the same sensor under near-identical conditions.
- SSSR router is broken: Use
--routing forest. See above.
Why spectral context?
There's a persistent problem in hyperspectral imaging that I don't see talked about much directly: a model trained on one scene generally can't be used on another.
You train on Indian Pines and get 97% accuracy. You run the same model on Salinas Valley and it falls apart — not because the materials are different (both scenes have vegetation, bare soil, crops), but because the model learned where things tend to be, not what they are spectrally.
Current HSI methods, including v0.1.x of this project, rely heavily on spatial context. A U-Net learns that "pixels in this area tend to be corn because they're spatially clustered with other corn pixels." That pattern holds within one scene. It means nothing in the next.
What should generalize is the chemistry. Chlorophyll absorbs light at around 680 nm because of its molecular structure — that's true whether you're looking at a corn field in Indiana or a rice paddy in Vietnam. Hemoglobin absorbs differently from collagen, regardless of which patient, which hospital, which staining protocol. Spectral fingerprints are determined by molecular bonds, and molecular bonds don't change between scenes.
If a model learned to classify from spectral evidence alone — no location, no neighbors — you could train it on two scenes and use it on fifty more. That's the goal of v0.2.x: remove all spatial context from the classification loop and force the model to rely entirely on the spectrum of each individual pixel.
I want to be upfront that this is an experiment. Whether the spectral-only approach delivers on that promise is something that needs to be demonstrated, not just argued. The v0.2.x evaluation on multi-scene datasets is what will actually tell us. But the reasoning seems sound, and it's worth trying — especially since I haven't found a clean, accessible implementation of this idea elsewhere in the literature.
What makes it difficult ?
Spectral variance within groups is often larger than between groups. What this means is that if element A has absorption characteristics that vary significantly within its group, it can be difficult to distinguish it from element B, even if B has different absorption characteristics overall. If A has features between 100nm and 200nm while B has features between 150nm and 250nm, then the region between 150–200nm becomes highly ambiguous.
Beyond overlapping signatures, real-world conditions rarely give us pristine data. Variables like illumination angles, shadows, canopy geometry, and atmospheric scattering constantly distort the readings. Two identical crop fields can yield distinct spectral profiles simply because one was imaged at noon and the other at 4 PM under thin clouds.
Because of these ambiguities, current methods in the literature—just like v0.1.x of this project—take the easy way out: they lean heavily on spatial context. They operate on the logical assumption that if 10 pixels around pixel X are corn, then X is highly likely to be corn as well. Spatial architectures like U-Nets excel at learning these neighborhood patterns, effectively smoothing over the spectral noise.
But this spatial crutch masks the underlying problem. It ties the model's performance to the specific spatial layout of the training scene. When we remove that spatial safety net in v0.2.x, the model loses the ability to infer a pixel's identity from its neighbors. It is forced to confront the raw, messy chemical reality of "mixed pixels" and atmospheric distortion on its own.
Tackling these hurdles—intraclass variance, atmospheric distortion, and high dimensionality—without relying on spatial crutches is what makes v0.2.x a formidable challenge. The model has to learn the true, underlying chemical reality amidst a sea of noise. Convex Hull Continuum Removal and SPT have proven to be quite effective at learning this spectral context, but further studies are needed.
What has been achieved so far ?
The incomplete v0.2.x build has been tested on Indian Pines under a strict spectral-only regime — 100 pixels per class for training, with 40% of available pixels used for smaller classes, and zero spatial context at any point in the pipeline. The result was 92% OA. No neighbors. No U-Net. No spatial smoothing of any kind.
That number alone isn't what's interesting. What's interesting is the per-class mIoU distribution. Across nearly all classes, mIoU sits consistently around 0.7 — the model isn't carrying a few easy classes while failing silently on hard ones. The only meaningful outlier is Class 7 (Grass-pasture-mowed), which has very few labeled pixels in the Indian Pines ground truth to begin with — a data problem, not a model problem. The uniformity across the remaining classes is the first real evidence that the model is learning spectral chemistry rather than spatial habit.
This result is preliminary. The full convex hull continuum removal implementation is not fully optimized — the simpler linear continuum removal currently outperforms it, which suggests the hull computation is introducing noise rather than removing it. That needs to be resolved before any result involving continuum removal can be reported cleanly. GPU utilization is also unoptimized, which makes thorough hyperparameter sweeps expensive and slow.
The ablation studies have not been completed or documented to a publishable standard. Claiming 92% OA without showing what each component contributes — SPT, continuum removal, dilation rates, ensemble size — is a number without a story. Those studies are in progress and will be reported in full once they are. Until then, treat this as a proof of direction, not a finished result: spectral-only classification on Indian Pines is viable, and the per-class consistency suggests the architecture is learning the right thing.
I will open source the code only once ghost-hsi v0.2.x has reached a stable release, achieved better numbers and has publishable ablation studies. For more details, feel free to reach out to me at IshuIsAwake.
Start here.
Install.
Install from PyPI. Indian Pines sample data is bundled — run ghost demo to get the file paths.
pip install ghost-hsi
ghost demoTrain.
Run the full GHOST pipeline with SPT. Use the paths printed by ghost demo.
ghost train_spt \
--data /path/to/Indian_pines_corrected.mat \
--gt /path/to/Indian_pines_gt.mat \
--loss dice --routing forest \
--base_filters 32 --num_filters 8 \
--ensembles 5 --leaf_ensembles 3 \
--epochs 400 --patience 50 --min_epochs 40 \
--out-dir runs/my_experimentPredict.
Run inference on the test split and compute metrics.
ghost predict \
--data /path/to/Indian_pines_corrected.mat \
--gt /path/to/Indian_pines_gt.mat \
--model runs/my_experiment/spt_models.pkl \
--routing forest --out-dir runs/my_experimentVisualize.
Generate a 3-panel segmentation figure: false colour composite, ground truth, GHOST prediction.
ghost visualize \
--data /path/to/Indian_pines_corrected.mat \
--gt /path/to/Indian_pines_gt.mat \
--model runs/my_experiment/spt_models.pkl \
--dataset indian_pines --routing forest \
--out-dir runs/my_experiment